Human 3’UTRome V1
![]() 1,461 Human 3'UTR clones ready for the 3’LIFE screen are now available HERE
1,461 Human 3'UTR clones ready for the 3’LIFE screen are now available HERE
please cite:
Kotagama K, Babb CS, Wolter JM, Murphy RP and Mangone M.
A human 3'UTR clone collection to study post-transcriptional gene regulation.
BMC Genomics. 2015 Dec 9;16:1036. doi: 10.1186/s12864-015-2238-1.
We have learned a lot from worms about 3'UTR RNA biology. Having the full 3'UTRome allows us to study PAS elements, miRNA targets, and alternative polyadenylation with a precision unthinkable a few years ago.
In addition to the sequences, we previously developed a genomic 3'RACE pipeline and cloned at least one 3'UTR isoform for every protein-coding gene (Mangone et al., 2011). This library was the first for a metazoan organism, and it allows for the first time the investigation of post-transcriptional gene regulation in vivo using high-throughput downstream analyses.
In human, unfortunately, we still do not have a complete cloned 3'UTRome. The extent of alternative polyadenylation is still largely unknown but estimated to occur in at least half of the transcriptomes. The majority of the data available comes from the re-annotation of ESTs and other genomic approaches, but nobody has tackled this question systematically, yet, using a multi-pronged approach as we did in worms.
We have now started the preparation of a human 3'UTRome library, cloning 3'UTR genomic regions for each of the ~20K human protein coding genes. When it will be completed, the human 3'UTRome clone collection will represent an incredible milestone in genome biology. It will provide an invaluable tool to study transcriptional termination, probe miRNA biology, study RNA binding protein elements and post-transcriptional gene regulation, and to better understand the biology of these gene portions in human disease. The human 3'UTRome resource will be publically available to the community, and will provide the foundation for further analysis of 3'UTR involvement in the broad range of human disease that they have been implicated in, including diabetes, Alzheimer's and cancer.
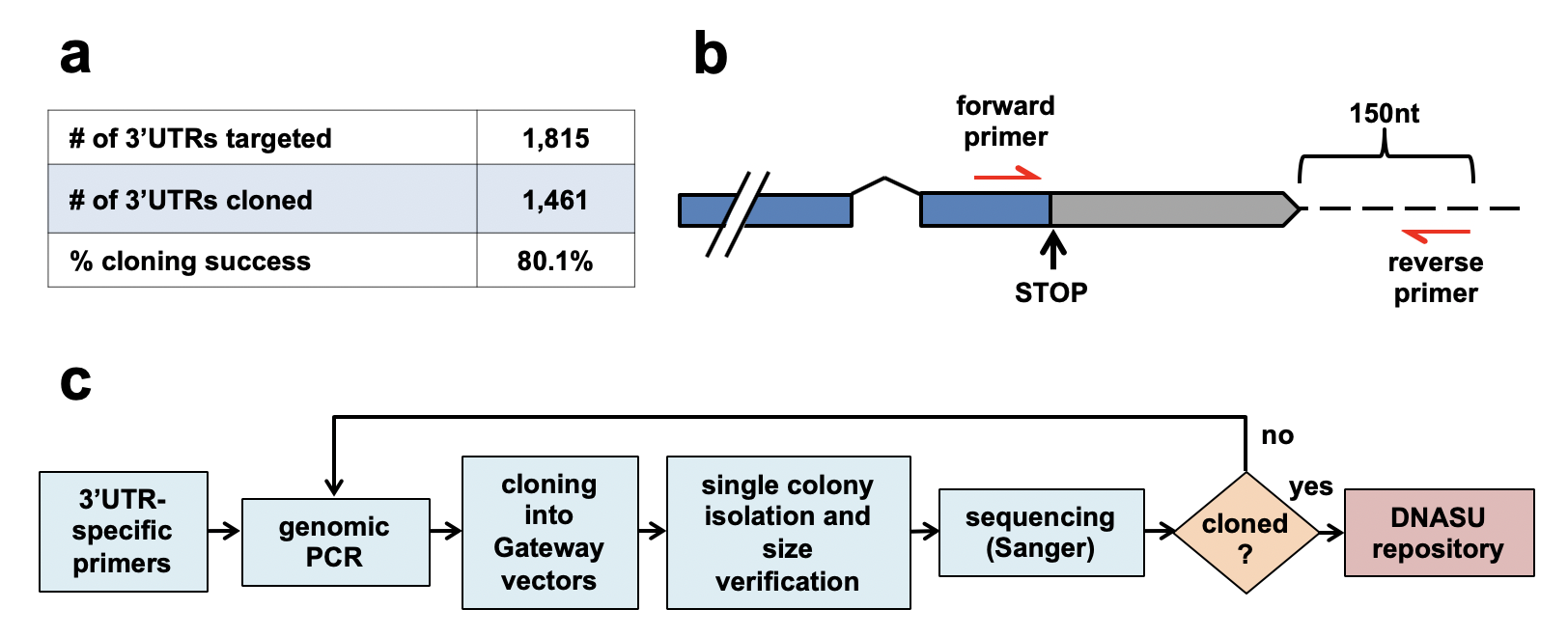
The human 3′UTRome cloning pipeline. We targeted a panel of 1,815 unique human 3′UTRs and successfully cloned and sequence verified 1,461 unique 3′UTRs (80.1 % cloning success). The forward primers used to amplify 3′UTR genomic loci were anchored within the last exon of each transcript, ending with the gene specific STOP codons. The reverse primers bound 150 nucleotides downstream of the annotated transcript. c Flow chart summarizing the cloning pipeline of the h3′UTRome v1. Genomic PCR was performed using 3′UTR specific primers and the PCR products were shuttled into Gateway® Entry vectors by recombinational cloning. Single cloned colonies were isolated and screened based on the expected 3′UTR length using PCR and gel electrophoresis. Bacterial colonies passing the screen were then re-arrayed, and the cloned 3′UTRs were sequenced using Sanger sequencing method. The sequence verified 3′UTRs were then submitted to the DNASU plasmid repository for public distribution. 3′UTRs that were not successfully cloned were subject to a second pass of cloning.
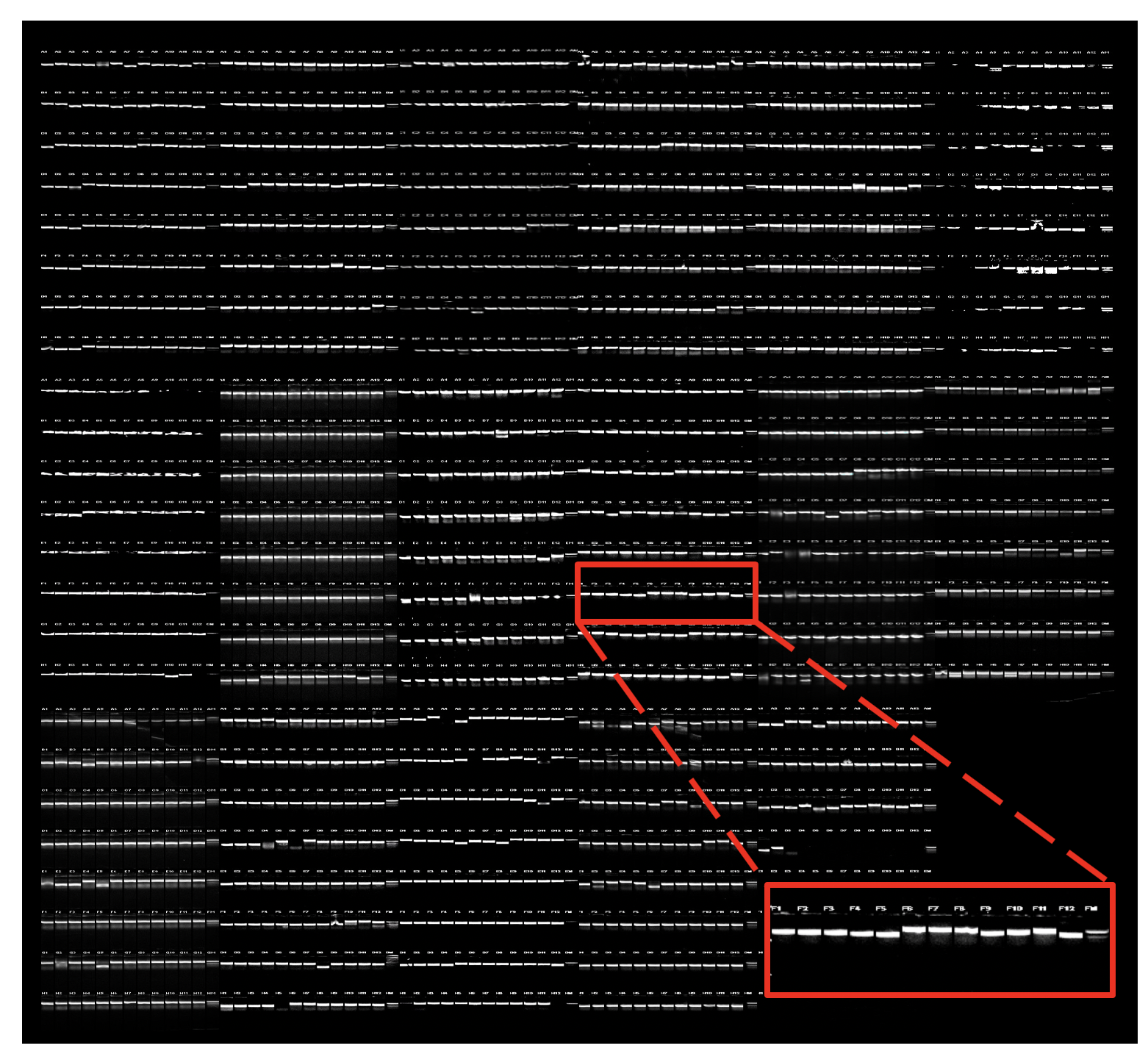

Electrophoretic analysis of PCR products from the complete h3′UTRome v1. The sizes of 3′UTRs from 1,461 PCR reactions were analyzed on ethidium bromide stained gels
